The language of life has limited characters. Just 4 letters A, T, G, and C (short for adenine, thymine, guanine, and cytosine) make up the genetic material in every organism. Of course, U (uracil) replaces T in the RNA. It is not too hard to imagine the pre-biotic soup consisting of many such molecules, which could have been the contenders to carry genetic information. DNA would have been selected among these for obvious reasons.
Many xeno-nucleotides (XNAs) have been synthesized in the lab and shown to be capable of heredity and evolution. There are also PNAs (peptide nucleic acids), which have proteins in their backbones and are heralded as a boon to nano-medicine. Why, then, nature settled on just 4 nucleotides? One thing that we can be sure of is that an odd number of nucleotides is not feasible due to complementary base pairing.
“If you want to understand life, think not of throbbing gels, think of information technology”
Richard Dawkins
Possible Nucleotides
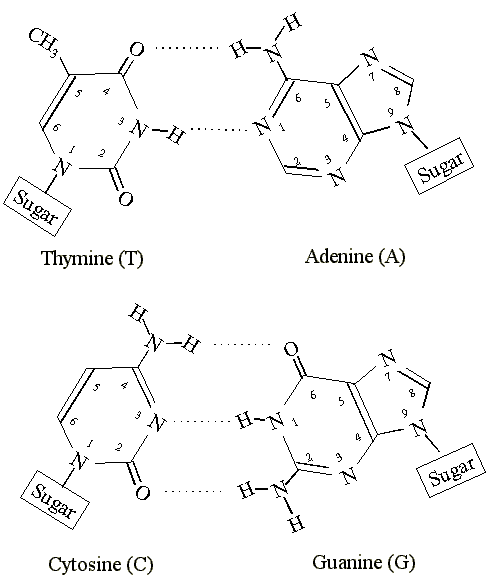
Donall Mac Donaill, an Irish scientist, came up with a paper in 2002 that was too obvious to have been ignored this long. He represented each of the nucleotides in 4 bits. The first three bits represented the possible interactions a nucleotide can have with its complementary nucleotide and the fourth bit represented the ring.
The hydrogen donating interactions were represented as 1 and accepting as 0. The pyrimidines got a 0 and the purines a 1. The nucleotides T, C, G, A were, hence, denoted as 0101, 1001, 0110, and 1010 respectively. The bits were also in accordance with complementary base pairing (for example A=T is 1010=0101). But, for 4 bits a nucleotide, there exist 24 = 16 possible nucleotides. These are also chemically possible.
Adding A Parity Bit
Luckily, Donaill had a background in computers. He knew about a system developed in the Bell Labs in the ’50s which added to an extra bit to the end of each signal to make the sum of digits either odd or even, as agreed upon earlier by the sender and the receiver. Any error in the signal changes the sum of the signal and, hence, the receiver knows something has gone wrong.
For instance, let this be the signal the sender intends to transmit
001 110 101 010 111 011
Adding a parity bit to make each signal subset evenly summed makes it
0011 1100 1010 0101 1111 0110
Any deviation from this and the receiver knows that there is an error. This system is employed in nearly everything from credit cards to online trading. Donaill argued that nature had a similar check for nucleotides.
Why Would Nature Prefer More Nucleotide Pairs
Greater the number of nucleotides, greater would be the diversity because of more permutations and combinations in the DNA sequence. But, it also makes it challenging for the cells to manage relatively equal concentrations of each nucleotide.
I know the nucleotides aren’t present in equal amounts in a DNA sequence, but the cell doesn’t know that and it must keep relatively equal amounts of each nucleotide at disposal.
Suppose there were only two nucleotides. They would be denoted as 0 and 1 and their relative concentrations would be equal and half of the total. To process DNA at any stage – replication, transcription, or translation – the sequence is run linearly.
The processing speed is then
1 bit X 0.5 relative concentration = 0.5 bit per unit time
For the naturally occurring scenario, each nucleotide can be denoted by 2 bits and has a relative concentration of one-fourth of the total. Thus, the processing speed is
2 bits X 1/4 relative concentration = 0.5 bit per unit time
For 6 nucleotides, the processing speed would be
log26 X 1/6 relative concentration = 0.43 bit per unit time
For 8 nucleotides, the processing speed would be
log28 X 1/8 relative concentration = 0.37 bit per unit time
Note that as the number of nucleotides increases, the processing speed drops gradually. The best speed is when there are either 2 or 4 nucleotides. Of these two cases, 4 nucleotides accommodate greater diversity.
A Fine Balance
The balance between processing speed and possible variation seems to be the plausible reason why nature settled on just four nucleotides. I expressed the processing speed per unit time and did not say anything about how much the unit was because that varies across species and the process concerned.
This is something like Mendel’s factors in the late 1800s, something deducted based on pure mathematical reasoning with physical proof yet to be validated.
Fun fact: at 2 bits per nucleotide, the entire human nuclear genome (with 3.2 billion base pairs) can be coded in just 756 MB of computer memory.
Read more here
Donaill, D. M., A parity code interpretation of nucleotide alphabet composition, Chemical Communications (Cambridge), Sep 21,2002
Why does DNA only use four nucleotides, Carl Brannen’s blog

